Directly read from a Pandas DataFrame
You can read directly from a Pandas DataFrame. Just give the features/predictors' column names as a list and the target column name as a string to the fit_dataframe method.
At this point, only numerical features/targets are supported but in future releases we will support categorical variables too.
<... obtain a Pandas DataFrame by some processing>
df = pd.DataFrame(...)
feature_cols = ['X1','X2','X3']
target_col = 'output'
model = mlr()
model.fit_dataframe(X=feature_cols,y = target_col,dataframe=df)
Metrics
So far, it looks similar to the linear regression estimator of Scikit-Learn, doesn't it?
Here comes the difference,
Print all kinds of regression model metrics, one by one,
print ("R-squared: ",model.r_squared())
print ("Adjusted R-squared: ",model.adj_r_squared())
print("MSE: ",model.mse())
>> R-squared: 0.8344327025902752
Adjusted R-squared: 0.8100845706182569
MSE: 72.2107655649954
Or, print all the metrics at once!
model.print_metrics()
>> sse: 2888.4306
sst: 17445.6591
mse: 72.2108
r^2: 0.8344
adj_r^2: 0.8101
AIC: 296.6986
BIC: 306.8319
Correlation matrix, heatmap, covariance
We can build the correlation matrix right after ingesting the data. This matrix gives us an indication how much multicollinearity is present among the features/predictors.
Correlation matrix
model.ingest_data(X,y)
model.corrcoef()
>> array([[ 1. , 0.18424447, -0.00207883, 0.144186 , 0.08678109],
[ 0.18424447, 1. , -0.08098705, -0.05782733, 0.19119872],
[-0.00207883, -0.08098705, 1. , 0.03602977, -0.17560097],
[ 0.144186 , -0.05782733, 0.03602977, 1. , 0.05216212],
[ 0.08678109, 0.19119872, -0.17560097, 0.05216212, 1. ]])
Covariance
model.covar()
>> array([[10.28752086, 1.51237819, -0.01770701, 1.47414685, 0.79121778],
[ 1.51237819, 6.54969628, -0.5504233 , -0.47174359, 1.39094876],
[-0.01770701, -0.5504233 , 7.05247111, 0.30499622, -1.32560195],
[ 1.47414685, -0.47174359, 0.30499622, 10.16072256, 0.47264283],
[ 0.79121778, 1.39094876, -1.32560195, 0.47264283, 8.08036806]])
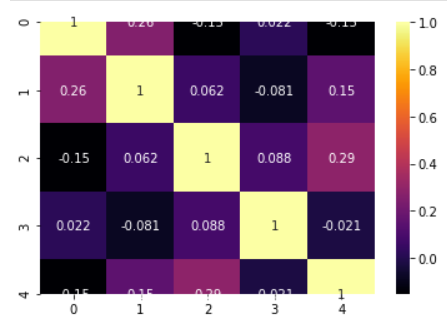
Correlation heatmap
model.corrplot(cmap='inferno',annot=True)
Statistical inference
Perform the F-test of overall significance
It retunrs the F-statistic and the p-value of the test.
If the p-value is a small number you can reject the Null hypothesis that all the regression coefficient is zero. That means a small p-value (generally < 0.01) indicates that the overall regression is statistically significant.
model.ftest()
>> (34.270912591948814, 2.3986657277649282e-12)
How about p-values, t-test statistics, and standard errors of the coefficients?
Standard errors and corresponding t-tests give us the p-values for each regression coefficient, which tells us whether that particular coefficient is statistically significant or not (based on the given data).
print("P-values:",model.pvalues())
print("t-test values:",model.tvalues())
print("Standard errors:",model.std_err())
>> P-values: [8.33674608e-01 3.27039586e-03 3.80572234e-05 2.59322037e-01 9.95094748e-11 2.82226752e-06]
t-test values: [ 0.21161008 3.1641696 -4.73263963 1.14716519 9.18010412 -5.60342256]
Standard errors: [5.69360847 0.47462621 0.59980706 0.56580141 0.47081187 0.5381103 ]
Confidence intervals
model.conf_int()
>> array([[-10.36597959, 12.77562953],
[ 0.53724132, 2.46635435],
[ -4.05762528, -1.61971606],
[ -0.50077913, 1.79891449],
[ 3.36529718, 5.27890687],
[ -4.10883113, -1.92168771]])
Visual analysis of the residuals
Residual analysis is crucial to check the assumptions of a linear regression model. mlr helps you check those assumption easily by providing straight-forward visual analytis methods for the residuals.
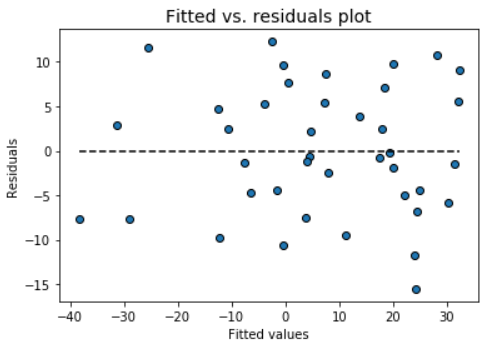
Fitted vs. residuals plot
Check the assumption of constant variance and uncorrelated features (independence) with this plot
model.fitted_vs_residual()
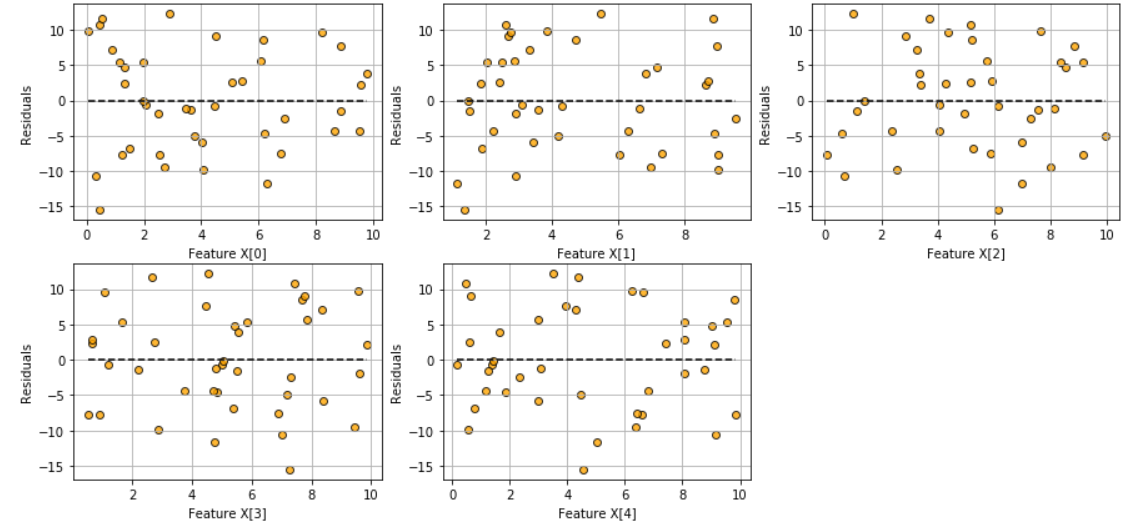
Fitted vs features plot
Check the assumption of linearity with this plot
model.fitted_vs_features()
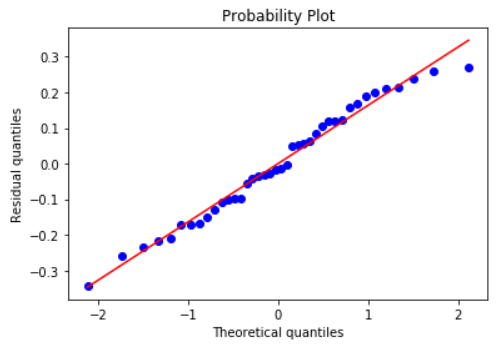
Histogram and Q-Q plot of standardized residuals
Check the normality assumption of the error terms using these plots,
model.histogram_resid()
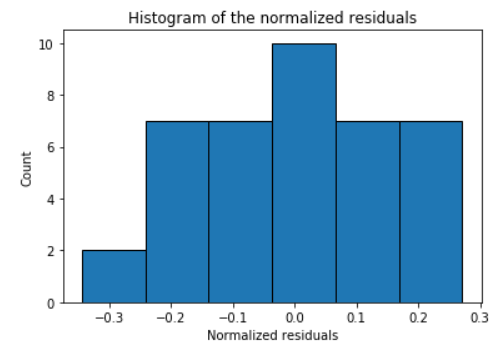
model.qqplot_resid()